Heart Failure (HF) is a progressive condition wherein the heart is unable to fill its chambers and/or pump sufficient blood into the arteries. HF is often secondary to cardiovascular disease (e.g., coronary artery disease, hypertension). Common symptoms include shortness of breath and reduced exercise capacity, physical findings (lung fluid and ankle swelling), and evidence of abnormal ventricular function. Many studies have identified biomarkers of HF that reflect various aspects of the heart failure syndrome such as oxidative stress, myocardial stretch, myocyte injury, matrix remodeling, neurohormonal activation, etc. Although many biomarkers have been postulated and evaluated, only brain natriuretic peptide (BNP) is routinely used in clinical trials for HF. Single molecule biomarkers have been useful in elucidating particular aspects of the underlying causes of HF. However, it is unlikely that any given biomarker can capture the complexity of the underlying disease processes. In the current era, an integrated, unbiased approach is possible, allowing investigators to discover, replicate and validate novel diagnostic and prognostic biomarkers for HF, and those that reflect responsiveness to therapeutic intervention.
Systems science moves beyond the study of single entities to entire systems, embracing the complex interactions between entities that make up these systems, and the interactions between systems and the environment, and constantly changing properties of these systems. Systems biology is the study of biological systems through modelling of the complex interactions across various levels within these systems (molecules, cells, tissues, organs, organisms). This type of interrogation is commonly achieved using molecular networks of interacting molecules where the interactions are based on curated databases (e.g., protein-protein interactions) and empirical data (e.g., using high throughput ‘omics’ platforms). Technological advances coupled with decreasing costs of obtaining high throughput ‘omics’ datasets have made it possible to obtain multiple sources of information for the same set of individuals, allowing in turn for a comprehensive understanding of heterogeneous diseases. A systems biological approach leverages the signal across multiple biological domains in order to explain the variability in the associated phenotype (Figure 1, inner). The combination of information across different biological layers helps unravel the complexities of the underlying disease processes. Biological and molecular processes are comprised of interactions between different biological layers such as the genome, methylome, transcriptome, proteome and metabolome. These interactions are often missed when each omic level is studied in isolation, leading to an increased number of false positives, loss of information and unreliable findings.
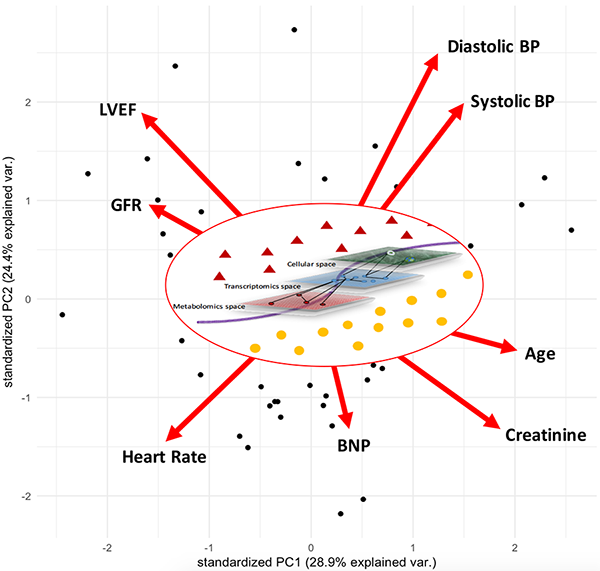
Given the multifactorial nature of HF, it is ideally suited for interrogation by systems biology approaches. Unsupervised learning approaches, which do not rely on prior phenotypic labels have been used to identify novel disease phenotypes and even improve phenotypic characterization. Figure 1 (outer) depicts a clustering of patients (black dots) based on clinical variables (red arrows) that are used for the diagnosis of HF. Each arrow is centered at zero, which refers to the average value of that variable, e.g., an individual at the origin has a clinical profile that corresponds to the average value of each of the clinical variables in the figure. Moving along the axes for any given variable corresponding to increases from the average value of that clinical variable. For example, individuals (depicted by the black dots) near the bottom of Figure 1 (along the y-axis) have an increased heart rate and levels of BNP compared to individuals at the top of Figure 1. Further, variables that are closer together (where the angle (θ) between the arrows is less than 90) are positively correlated whereas variables that are further apart (90 < θ ≤ 180) are negatively correlated. These approaches allow the simultaneous interrogation of many variables for the characterization of patients and can be used to complement and enhance medical decision making.
My postdoctoral fellowship at the Prevention of Organ Failure (PROOF) Centre of Excellence combines clinical, molecular, computational and statistical disciplines in order to develop a deeper understanding of HF phenotypes and their underlying mechanisms. Specifically, with my team members, I am developing blood-based biomarker tests that differentiate HF patients with reduced ejection fraction (baggy hearts) as compared to preserved ejection fraction (stiff hearts). Further, I am interrogating the molecular mechanisms of HF patients with improved as compared to deteriorated heart function over time. Lastly, I am developing novel methodologies for the integration of multiple omic datasets for the classification of HF phenotypes. Earlier recognition and treatment of HF will significantly reduce 1) the burden of future HF symptoms for individual patients, 2) the frequency of HF hospitalizations, and 3) the risk of premature death while 4) improving quality of life and productive employment. If we integrate our systems biological results with social, ethnic, and environmental factors we will ultimately be able to represent risk and disease course in real-time by harnessing the power of systems science.